Enzyme engineering with evolution
Ever since the very first living thing emerged on Earth, evolution has shaped the development and diversity of life. The drive to survive and thrive has resulted in a spectacular range of living things that seem like perfect fits for their particular environmental niches, as favourable genetic variants are passed down from one generation to another.
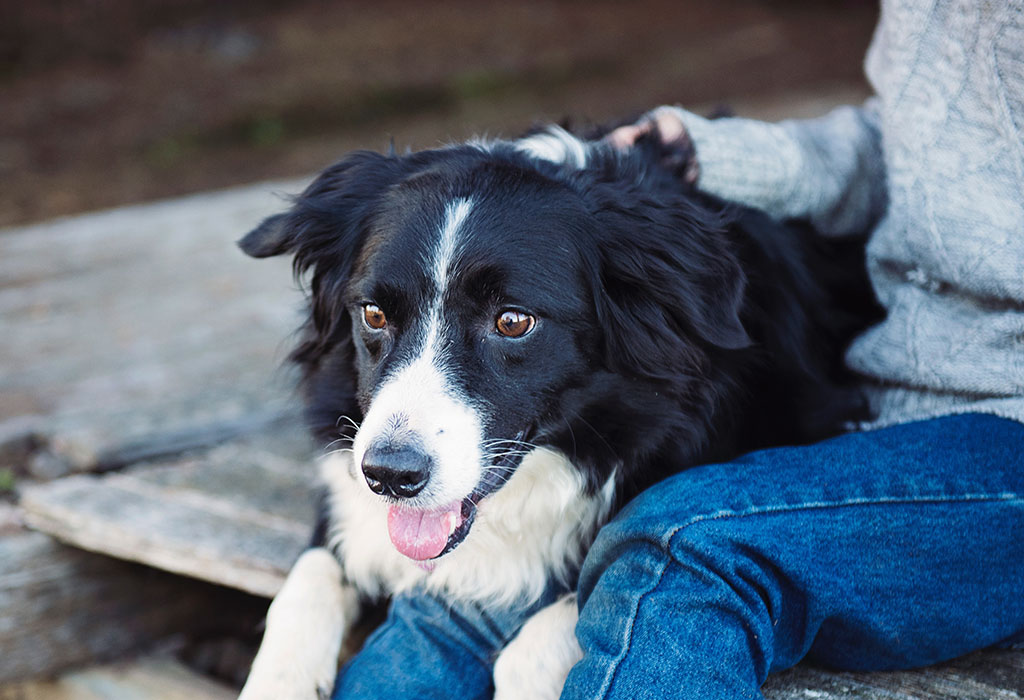
Humans have been tinkering with the tools of life for thousands of years, ever since we first started deliberately breeding plants or animals to bring out the traits we wanted—a tree with sweeter fruit, perhaps, or dogs that are loyal and easy to train.

We’ve also been able to harness the power of evolution to create biochemical tools that can build better medicines and cleaner fuels. It’s a feat that won chemical engineer Frances Arnold the 2018 Nobel Prize for Chemistry for her work in directed evolution of enzymes (jointly shared with biochemists George Smith and Gregory Winter for a separate discovery).
Enzymes are super useful biological molecules that speed up (catalyse) the rate at which chemical reactions take place, without being used up in the reaction itself. They’re responsible for all sorts of essential functions, from respiration to digestion.
They’re also extremely complex. Several thousand ‘building blocks’ (mostly amino acids) can be involved, all folded up into precise, intricate 3D shapes. And shape is very important: enzymes only work when they can interact with the other molecules involved in a reaction, somewhat like puzzle pieces fitting together, so if the shape isn’t right, nothing happens.
This complexity means designing enzymes for specific purposes is very difficult. Those amino acids can be combined and folded in an almost infinite number of ways, and the slightest change can have significant impacts on how well they can do their job (if they still work properly at all).

In the early 1990s, scientists were taking a logical and systematic approach towards developing custom-built enzymes that had just the right properties and shapes to catalyse specific reactions. It was tedious, slow and unpredictable. Frances Arnold came up with another idea: why not rely on ‘survival of the fittest’ principles to build and select the right enzymes instead?
Arnold wanted to modify an enzyme called subtilisin, a protein-digesting enzyme found in bacteria. It normally only works in water-based environments, but Arnold wanted to modify it to catalyse reactions in an organic solvent instead. This is such a fundamentally different kind of environment that, normally, the enzyme’s shape would radically change and it wouldn’t work at all.
Instead of trying to predict the right architectural configuration for her enzyme from the start, Arnold started tinkering with the section of the bacteria’s genetic code that was responsible for making the enzyme, deliberately creating random mutations in the DNA. She then introduced these modified genes into some bacteria, which generated thousands of different variations of the enzyme.
The next challenge was to work out which of these variants were the best at doing their job in an organic solvent. Subtilisin is good at breaking down casein, a protein found in milk. Arnold created a culture medium containing both the organic solvent and casein, popped onto that medium the bacteria containing these new enzyme variants, and was able to see which variants were best at digesting the protein. The winners were selected and went on to compete in the next round of random mutation generation.
With each new generation, enzyme variants were produced that worked better and better in this strange new organic solvent environment. Arnold found a subtilisin variant that worked 256 times better in the organic solvent compared to the original enzyme after just three rounds of this process. She’d found her winner.
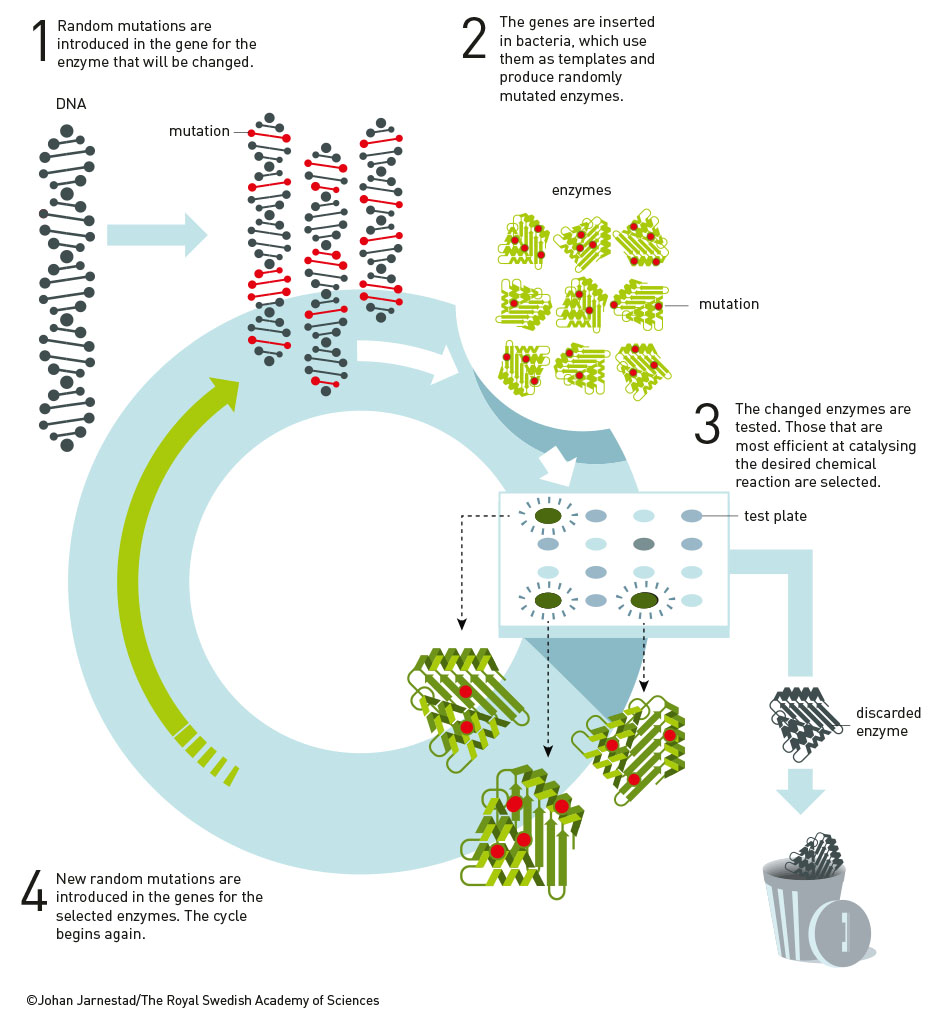
It would have been near-impossible for humans to predict exactly how they needed to modify subtilisin to achieve that desired result. With the evolution process doing the heavy lifting, Arnold was able to rapidly engineer an enzyme tailored for a specific task in a fraction of the time.
What’s more, with this technique, it is possible to create enzymes that wouldn’t otherwise exist in nature at all. Arnold’s lab successfully engineered enzymes that could make bonds between carbon and silicon atoms, which isn’t something that any living thing on Earth is able to do. These new enzymes could offer a cheaper, easier and more sustainable method for manufacturing products ranging from paints and sealants to medicines and medical devices—methods that currently rely on using lots of expensive metal-based catalysts.
Directed evolution of enzymes has also paved the way towards producing more environmentally-sustainable fuels by developing an enzyme that can turn simple sugars into a key component of biofuels. Arnold’s lab is part of a ‘green chemistry’ future where chemical reactions are faster, less wasteful and have lower environmental impacts. Using nature’s toolkit to create new ways of doing chemistry has utterly revolutionised what’s possible in laboratories around the world.